We have recently been working a lot on the characterization of our neurospheres. Below are the most interesting data from this.
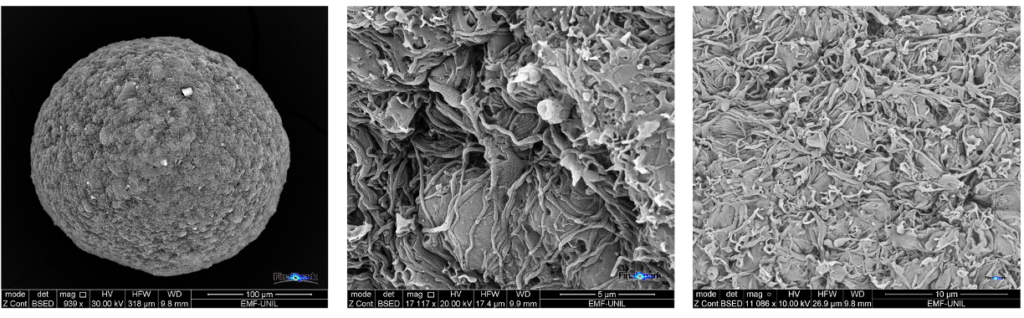
We acquired our first electron microscopy (ESEM-Quanta) images our neurospheres . Several samples were first stabilized in glutaraldehyde and then prepared for observation by our supplier. During several hours we had the chance to interactively explore the incredibly complex microstructure of our mini brains. The three images below show one entire neurosphere (left) and closeups of structures on the surface (middle and right), showing neurites and somas.
In addition to visual inspection, we also performed our first PCR!
This is a particularly critical step since we are differentiating neuro-progenitors in our own lab and it is important to verify exactly which type of cells we are generating.
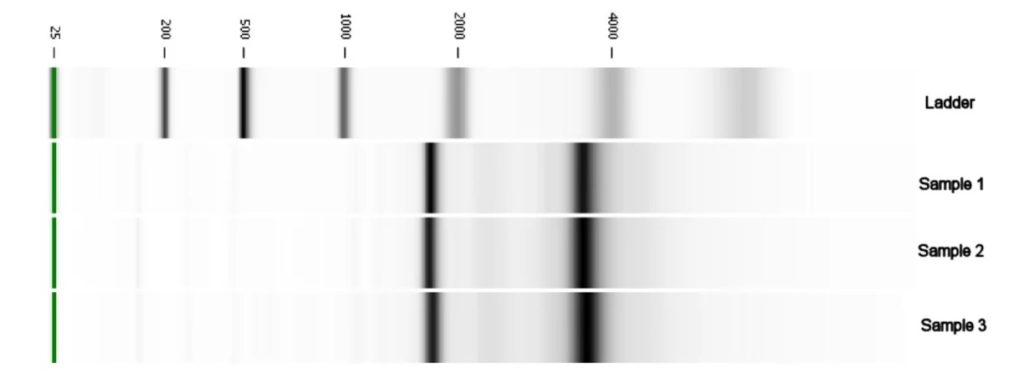
We performed an RNA extraction using 30 neurospheres split in 3 sets. RNA was first checked for quality using electrophoresis. The picture below shows the result for those samples (and the reference ladder on first line). We have the expected ribosomal 18S and 28S RNA bands for eukaryotes, both bands are well defined and the computation shows also a correct 2:1 ratio between their respective intensities. This confirms a successful RNA extraction.

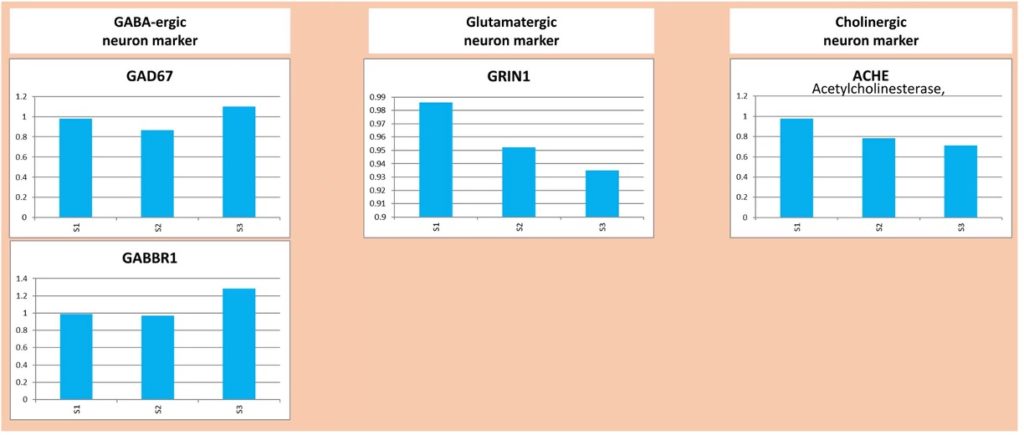
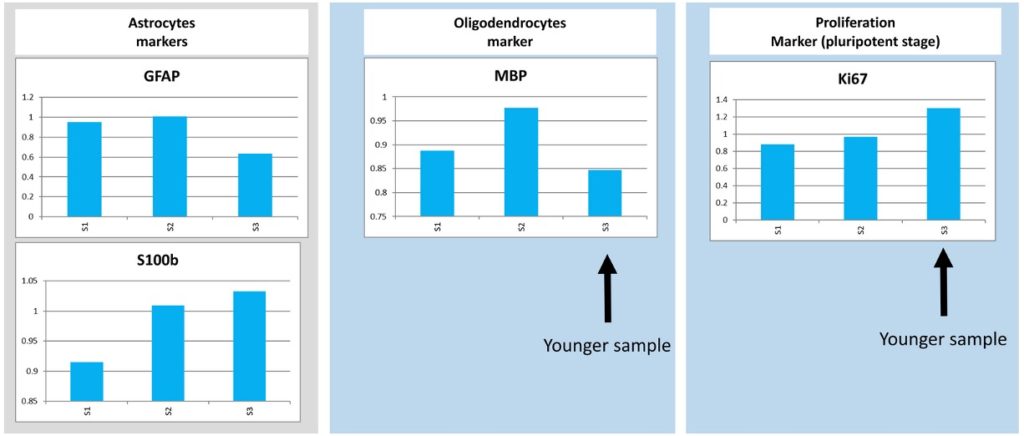
Our samples could then be analyzed by qRT-PCR. 14 different markers were used. The analysis of the results showed the presence of GABAergic, Glutamatergic and Cholinergic neurons. In addition to neurons, we also found glial cells like Astrocytes and Oligodendrocytes.
The conclusion was that our neurospheres seem to feature the expected cell types laying out the basis for a functional neural network!
We also continued our activity on electrophysiology. The core of our approach is based on interactions with the neurospheres using MEAs (Multi Electrode Array). In practice, this interaction is performed by stimulating electrically some neurons and measuring their responses. We already succeeded last year in eliciting reliable spike activity, however it required statistical analysis over seconds or minutes of activity.
We are proud to announce that we are now able to get reliable answers more than 10 times faster, typically within 50 milliseconds! This required a fundamental redesign of our hardware and software system.
The image below illustrates this behavior by showing:
- Horizontally, the experiment Id, each corresponding to 1 stimulation and a 100 ms recording of the response. In total, 600 stimulations have been performed every 6 seconds during one hour
- Vertically, the 100 ms recording showing the delay between answer and stimulation for each experiment (ms)
- Each dot corresponds to a spike
It can be noticed that the network responds extremely quickly to each stimulation (less than 20 ms), and the responses are very reliably (for 94% of the stimulations).
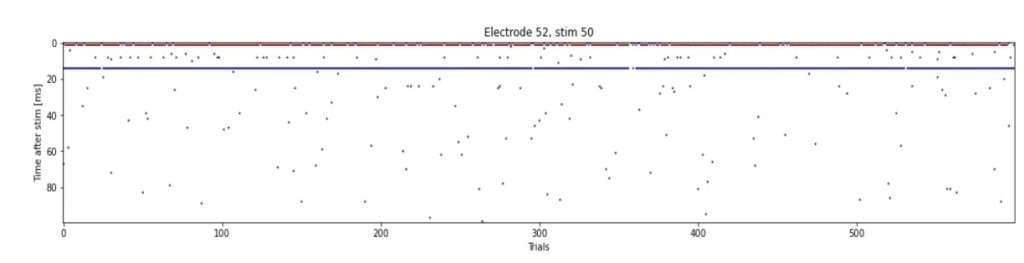
Moreover, by overlaying the 600 responses, we can confirm that this is a well-formed typical action potential recording:
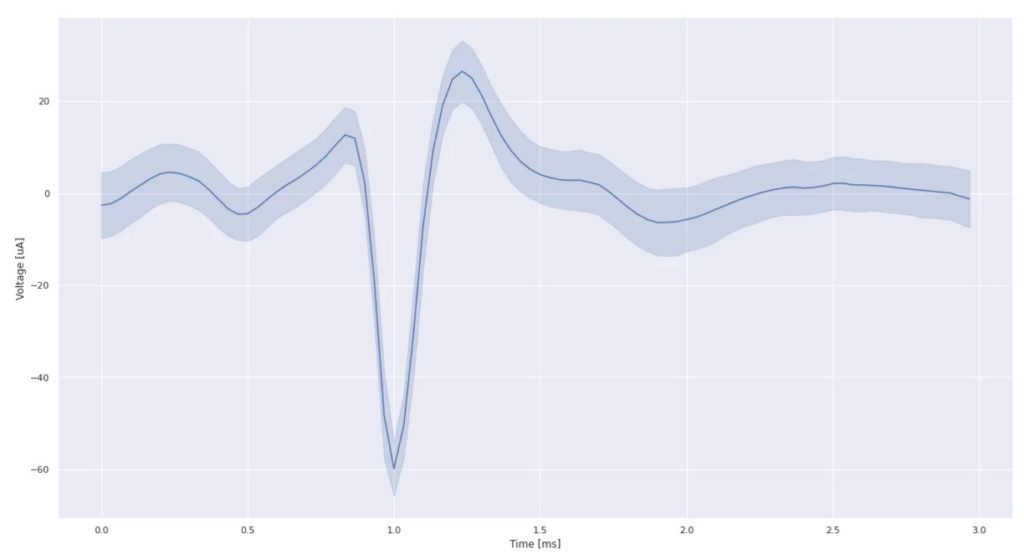
In parallel, we also worked on the microfluidics with two objectives in mind:
- Improve long-term reliability: After breaking our lifetime record of 120 days last year, we are now aiming to reach over 1 year of continuous recording of a single neurosphere. For such long durations, we need more durable pumps than our current syringe pushers and peristaltic pumps. For this purpose, we have started testing a new generation of mechanical pump since several weeks. The preliminary results (after many tunings) seem very promising but still need confirmation in the coming months.
- Chemical stimulations: The capability to inject specific molecules was planned already two years ago and will open an entire new set of possibilities, in particular for pharmacological tests. Indeed, the state of the art provides a handful of molecules that can be used to characterize the neurons and to validate the presence of various synaptic receptors. Such tests, when coupled with electrostimulation, also enable to provide information about the internal connectivity of the network (antidromic or synaptic connections for instance). These is precious information if we want to be able to train our networks to perform specific tasks, which remains our goal.
Experimental R&D platform fully operational
The 2nd and 3rd quarter of 2021 have been partly devoted to solving recurring issues which were drastically limiting our ability to do good science.
The first issue was related to the micro-fluidic setup where we faced multiple challenges. After exploring and combining different approaches we found ways to consistently avoid both bubbles (which kill a neurosphere in 15 minutes) and perfusion overflows (which take-off neurospheres from the electrodes). The final solution consists in a well-balanced combination of micro-fluidic pump technologies (peristaltic/syringe), time-dependent flow rate strategies, gravitational strategic positioning of Falcon 50, and tubing. The micro-fluidic setup now allows us to keep our neuronal cell cultures alive 24/7 without the need to manually take care of medium changes.
Our latest neurosphere lifetime record is an astonishing 87 days!
The second issue was electric noise on our electrodes which hindered us from performing consistent signal recording and analysis. The noise problem has been partially solved through rewiring a part of the AC power lines and optimizing the placement of the electrical equipment in the lab. As an example, the freezer which was relocated into a separate room at a fair distance of the lab.
We have now 4 MEA (Multi-Electrode Arrays) connected to the system allowing us to run simultaneous experiments on up to 16 neurospheres. This setup generates a substantial amount of data (100 GB/day). The data contains mainly voltage recordings of the 128 electrodes and pictures taken by the various microscopes from the lab and the incubator cameras.
In addition to the neurophysiological data, we also record 24/7 a myriad of environmental parameters like the pressure (lab+incubators), temperature, humidity, microfluidic flow rates, medium color, etc. The data is not only recorded, but also processed and stored 24/7 for analysis. Processed data includes for instance average spiking frequencies, timestamps, ISI (Inter Spike Interval), etc.
In order to use the data efficiently we have for 18 months been refining what can now be called a Bioelectronic Neuroscience Platform (BNP). The platform is basically an integrated set of connected web applications that provide an efficient way to manage and use all this wealth of information. Besides analytical tasks the platform allows to trigger automatic alarms (incubator door opening or temperature deviation for instance) and remotely control many of the day-to-day operations (taking pictures, pump control, remote activation of water bath pre-heating , …).
The diagram below shows all the applications that compose the BNP. In one single place it enables to do absolutely everything we need, for instance:
- Automatic computation and archiving of metrics for spiking activity analysi
- Camera application for monitoring and automatic analysis
- Micro-fluidic pumps programming or controlling (peristaltic and syringe )
- Real-time camera view for positioning of neurosphere in laminar flow hood
- Real-time display and archiving of environmental sensor data
- Graphic interface for direct access to time series database of measurements
- Control of microscopes for archiving and automatic analysis
- Automatic stimulation and analysis using integrated Python language
- Integrated lab notes system regarding all experiments and maintenance tasks
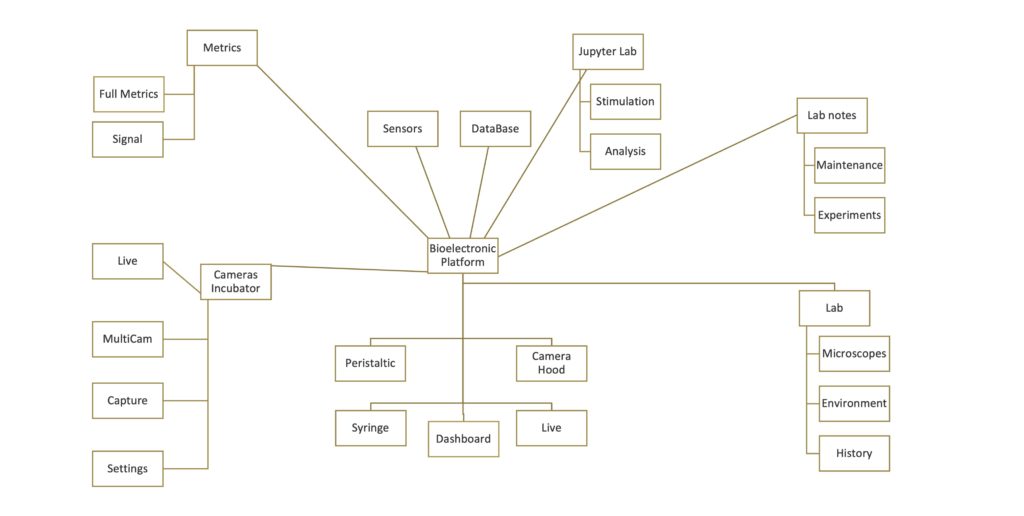
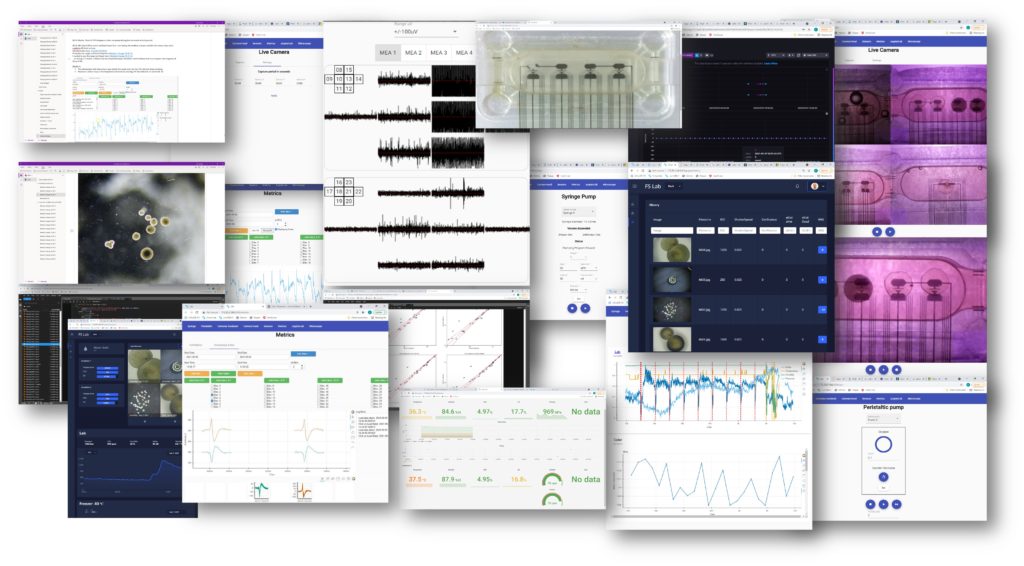
The image below shows a screenshot of each of the 20 web applications that constitutes our BNP.
Functionally the BNP can be divided into 4 categories:
- Display:like the real-time activity view, the dashboard or the camera hood.
- Programming: like the picture capture interval app or peristaltic pumping frequency and rate app.
- Interactive: like the metric computation or the microscope picture analysis by artificial neural networks (confluence, neurosphere counting, etc)
- Editing: like the tool for lab notes or python interface, those are the only applications that are not custom made: we use oneNote and JupyterLab.
On the more scientific, and less engineering, side, one of the important results that we obtained was the proof that we can reliably generate and record electrical responses to an electrical stimulation. While this observation may sound obvious given the state of the art in electrophysiology, it is for us of significant value because:
- There is only little state of the art regarding the electro-stimulation of hIPSC neurospheres
- We performed it in our own lab with our specific set of tools (knowing that the reproducibility of known results is not always given for granted in biology…) on long living neurospheres
- The electrical answer to the stimuli were strong, statistically reliable and reproducible
For illustrative purposes, the diagram below summaries one experiment on a specific electrode connected to a neurosphere:
- The horizontal axes represents the time with a total of 15 seconds of recording time
- each small vertical line represents a time stamp at which the electrode voltage exceeded 5 times the standard deviation of the signal (computed on a sliding window of 3 seconds)
- each row represents a different stimulation experiment (60 stimulations were performed here)
- the vertical red dashed line shows the time of stimulation (2 stimulations at 50mS interval of 100nA)
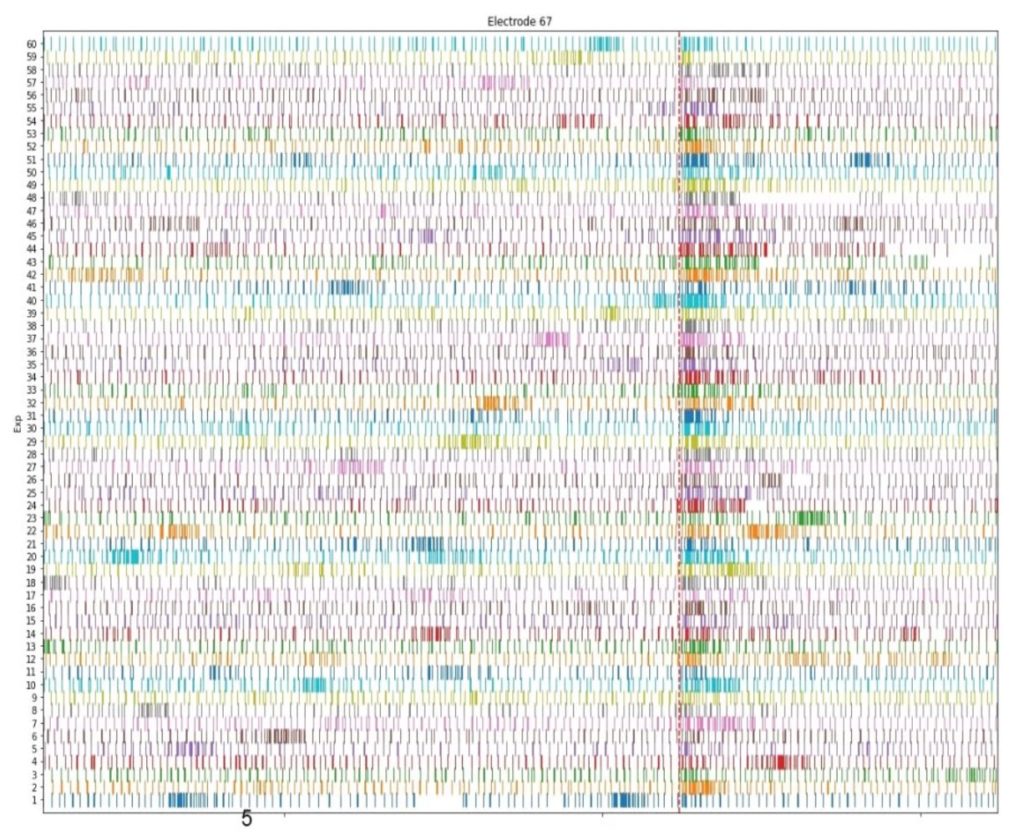

In the diagram we can observe that the stimulation clearly induces a statistically significant response. It is commonly believed that this type of response is a required condition for obtaining LTP (Long Term Potentiation), which remains our primary objective for the time being. As a next step we will reproduce the stimulation protocols described in literature (for example the works of Odawara and Maccione regarding hIPSC stimulations and Larson for theta-burst stimulations) and hope that it leads us to consistent and reproducible LTP.